3/24/2018 – Blog Post
Eleanor Lin
During the March 24th meeting, members were introduced to BLAST, the Basic Local Alignment Search Tool (https://blast.ncbi.nlm.nih.gov/Blast.cgi). The focus was on using Nucleotide BLAST to match RNA sequences from a file type called FASTA to RNA sequences in the NCBI database. FASTA files contain a name for each sequence and the order of the nucleotides in the sequence. Lines from a FASTA file can be copy-pasted into BLAST, which will provide a visual summary of the matches that it has found for the input sequence. It also lists out information for each individual sequence match.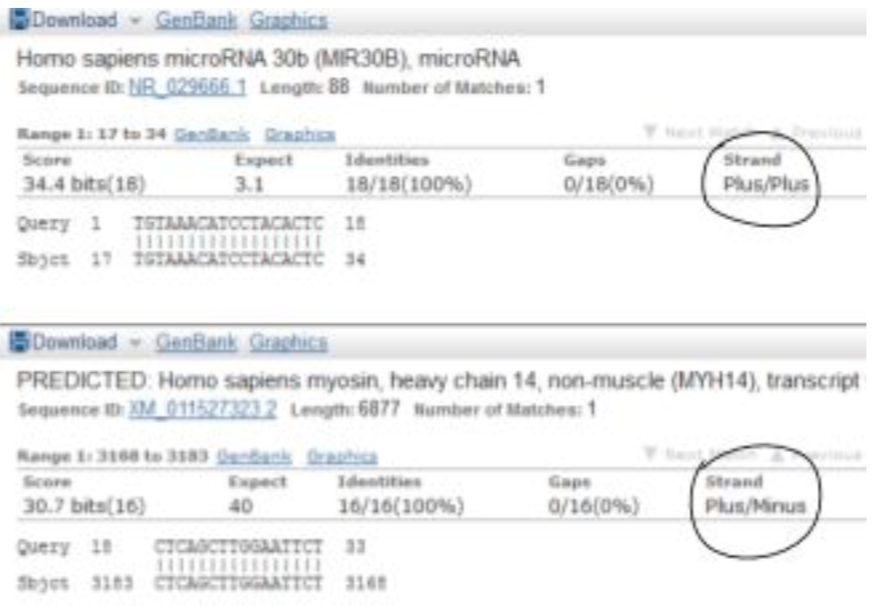 During the meeting, the concept of reverse complementary DNA sequences was reviewed, in order to explain why BLAST will label some sequence matches as “Plus/Plus” and others as “Plus/Minus.” DNA has two strands (it is a double helix) that go in opposite directions. One strand is designated the “positive” strand, and one is “negative.” DNA has a 5’ end and a 3’ end. When RNA is transcribed from the positive strand, it will be transcribed from the 3’ end towards the 5’ end of the DNA. The 5’ end of the RNA will match up with the 3’ end of the DNA, and vice versa. The RNA goes in the reverse direction compared to the DNA, but its base pairs still match (e.g. G to C). The reverse complementary RNA for a positive strand DNA sequence will be identical to the corresponding negative strand DNA sequence. For the example below, the reverse complementary RNA for the positive strand, read from the 5’ end, would be CAUCCU . . . the same as the negative strand, only with T’s replaced by U’s.
During the meeting, the concept of reverse complementary DNA sequences was reviewed, in order to explain why BLAST will label some sequence matches as “Plus/Plus” and others as “Plus/Minus.” DNA has two strands (it is a double helix) that go in opposite directions. One strand is designated the “positive” strand, and one is “negative.” DNA has a 5’ end and a 3’ end. When RNA is transcribed from the positive strand, it will be transcribed from the 3’ end towards the 5’ end of the DNA. The 5’ end of the RNA will match up with the 3’ end of the DNA, and vice versa. The RNA goes in the reverse direction compared to the DNA, but its base pairs still match (e.g. G to C). The reverse complementary RNA for a positive strand DNA sequence will be identical to the corresponding negative strand DNA sequence. For the example below, the reverse complementary RNA for the positive strand, read from the 5’ end, would be CAUCCU . . . the same as the negative strand, only with T’s replaced by U’s.
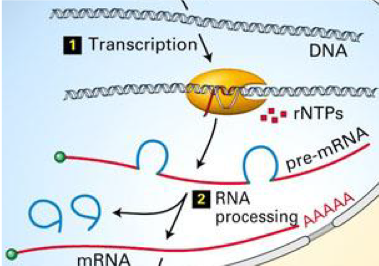
BLAST finds all the matches it knows, from both positive and negative strand DNA and RNA; this explains why some matches are Plus/Plus and others are Plus/Minus. However, another element of complexity is that RNA is processed by having parts removed, i.e. introns. So not all the RNA that is initially transcribed from the DNA makes it into the final RNA product. Depending on which introns are removed from the RNA, different “isoforms” (versions) can be created. The process of creating isoforms by processing the RNA differently is called “alternative splicing.” BLAST finds many different matches for the RNA sequence input that may be slightly different, or have gaps, due to alternative splicing.
Image: In number 2, “RNA processing,” the blue loops that are removed from the red pre-mRNA are the introns.
Image from miRcore summer camp slideshows.
The meeting concluded with suggestions from members on how miRcore leadership could be determined in the future, and what qualities a good miRcore leader should have. Members proposed a mix of the application and election processes in order to choose leaders, and qualifications such as experience, attendance, and, of course, compassion. Some commented that the list of good leadership qualities generated at the meeting included traits that were at odds with each other (being assertive versus being open-minded), but agreed that character was an important determining factor for good leadership. The discussion is still open, as miRcorers determine the direction of their organization in the years to come.