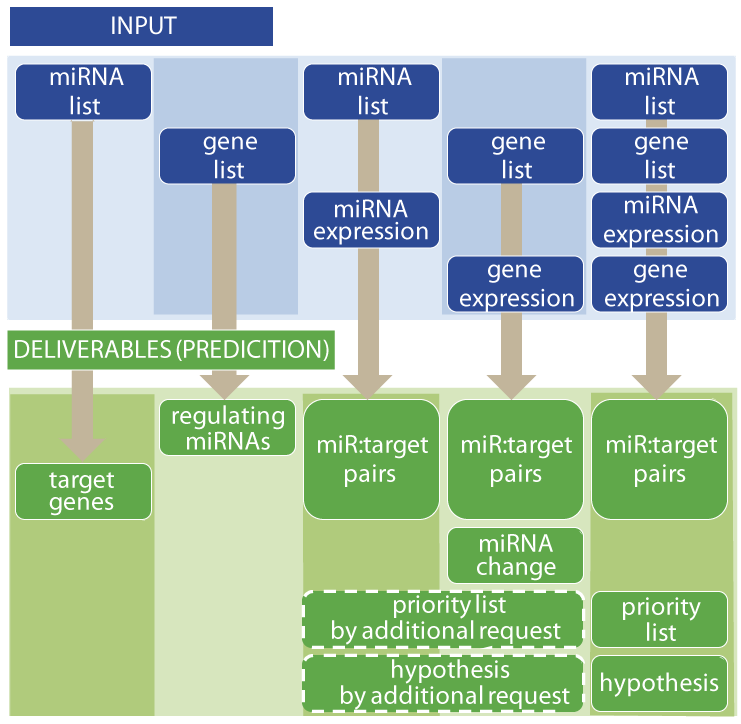
Deliverables
Efficient Research
In genetic research, priotizing which genes to study among tens of thousands is a daunting task. Our deliverables usually yield less than 2-10 fold fewer miR-target pairs to explore than do other conventional miRNA target prediction methods. Given expression data, we can further increase the specificity of biomarker lists. Given paired miRNA and gene expressions, our systems biology methods, including miBridge and mirFilter, usually yield fewer than 10 related RNAs without bias due to previous knowledge. Using gene expression alone, we identify potentially important miRNA function changes. We also provide hypotheses and priority lists for systems given same-sample miRNA-gene expression data. We ask for shared authorship should we significantly contribute to an intellectual advance.
miRNA Target Identification
miBridge (1) offers highly specific target gene predction, reducing time and cost in research. For example, miBridge successfully predicted that genes in the canonical Wnt signaling pathway were all targeted by miR-34, sparing researchers considerble effort (2).
1. July 2009, Genome Research, Vol. 19, No.7, pp.1175-1183.
2. November 2011, Science Signaling, Vol. 4, Issue 197, p. ra71.
Generate mirFilter Networks from Expression Data
mirFilter identifies differences between two conditions (e.g., metastatic and confined cancers, lithium-effective and no-lithium-effective groups) based on miRNA – gene expression correlations.
Small RNA Sequencing Analysis
Not only we are miRNA specialists, we are also proficient in tRF (tRNA-derived fragment) analysis, as seen in our work identifying functional tRFs with respiratory syncytial virus infection (3). You will receive high quality small RNA-seq analyzed data including both miRNAs and tRFs. Each dataset will be handled as collaborative scientific research and tailored to your needs at a cost comparable to other service providers. For large academic service cores, we may reduce your data process turnover time by complementing your expertise.
If you need small RNA sequencing experiments, we can also provide such services from total RNA in collaboration with the University of Texas Medical Branch. Additional services such as Northern blot confirmation are also available with UTMB.
- February 2013, Molecular Therapy, Vol. 21, No.2, pp.368-379.
